Data Updates
November 26th, 2025
The HTAN DCC has finalized the seventh release of HTAN data. The release includes 25416 new files for 422 participants and 1536 biospecimens from several atlases:
- Level 1-3 Bulk DNASeq data for 102 participants and 216 biospecimens
- Level 1-3 Bulk RNASeq data for 92 participants and 412 biospecimens
- Level 2 CODEX data for 32 participants and 52 biospecimens
- Level 2-3 CyCIF data for 49 participants and 59 biospecimens
- Level 1-4 Electron Microscopy data for 12 participants and 23 biospecimens
- Level 2 H&E data for 115 participants and 268 biospecimens
- Level 2 IMC data for 16 participants and 24 biospecimens
- Level 2-4 Imaging data for 84 participants and 212 biospecimens
- Level 2 MxIF data for 102 participants and 177 biospecimens
- Level 2-3 NanoString GeoMX DSP data for 51 participants and 97 biospecimens
- Level 2-3 mIHC data for 24 participants and 50 biospecimens
- Level 1-2 scDNASeq data for 4 participants and 11 biospecimens
- Level 1-4 scRNASeq data for 157 participants and 267 biospecimens
June 9th, 2025
New data from BU, Stanford, and WUSTL is available as part of Release 6.3:
- 10X Visium data for 1 participant and 32 biospecimens
- Level 1 Bulk RNASeq data for 15 participants and 21 biospecimens
- Level 2 H&E data for 89 participants and 430 biospecimens
- Level 1-4 scDNASeq data for 2 participants and 159 biospecimens
May 15th, 2025
We updated publication data to correct some file linking errors, and to reflect latest publication status. We also fixed a data access issue related to mzML files.
April 16th, 2025
New data from BU, CHOP, Duke, MSK, Vanderbilt, and WUSTL is available as part of Release 6.2:
- Level 1-4 and auxiliary 10X Visium data for 2 participants and 2 biospecimens
- Level 1-3 Bulk DNASeq data for 17 participants and 22 biospecimens
- Level 1-3 Bulk RNASeq data for 56 participants and 176 biospecimens
- Level 2 CODEX data for 3 participants and 8 biospecimens
- Level 2 H&E data for 8 participants and 10 biospecimens
- Level 3-4 Imaging data for 2 participants and 4 biospecimens
- Level 1-4 scRNASeq data for 48 participants and 91 biospecimens
March 19, 2025
Additional 25K Level 1-2 imaging and sequencing files are now accessible via CRDC-GC!
December 11th, 2024
We fixed an issue with manifest generation for CRDC-GC. This previously resulted in some files showing up in CRDC-GC while they weren't available yet.
November 22nd, 2024
More than 30K Level 1-2 imaging and sequencing files are now accessible via CRDC-GC!
November 5th, 2024
New data from BU, MSK, and Vanderbilt is available as part of Release 6.1:
- Level 1-2 Bulk DNASeq for 63 participants and 207 biospecimens
- Level 3 Bulk DNASeq for 49 participants and 66 biospecimens
- Level 1-4 scRNASeq data for 38 particpants and 91 biospecimens
- Level 2-4 Vectra and COMET imaging data for 24 participants and 98 biospecimens
August 26th, 2024
The HTAN DCC has finalized the sixth release of HTAN data. The release includes 43656 new files for 369 participants and 1926 biospecimens from several atlases:
- Level 1-4 and auxiliary 10X Visium data for 95 participants and 190 biospecimens
- Level 1-2 Bulk DNASeq data for 37 participants and 101 biospecimens
- Level 1-3 Bulk RNASeq data for 77 participants and 260 biospecimens
- Level 2 CODEX data for 41 participants and 56 biospecimens
- Level 2-4 CyCIF data for 20 participants and 119 biospecimens
- Level 3-4 Imaging data for 70 participants and 697 biospecimens
- Level 2 MERFISH data for 38 participants and 41 biospecimens
- Level 1-3 NanoString GeoMX DSP data for 50 participants and 276 biospecimens
- Level 1-3 Slide-seq data for 25 participants and 45 biospecimens
- Level 3 mIHC data for 4 participants and 22 biospecimens
- Level 1-3 sc/snATACSeq data for 18 participants and 24 biospecimens
- Level 1-4 sc/snRNASeq data for 103 participants and 123 biospecimens
- 10X Xenium ISS data for 1 participant and 1 biospecimen
- ExSEQ data for 9 participants and 9 biospecimens
June 3rd, 2024
New data from BU, and Stanford is available as part of Release 5.2:
- Level 3 HI-C-seq data for 10 new participants
- Level 1 sc/snRNASeq data for 14 new participants
April 23rd, 2024
New data from BU, HTAPP, CHOP, OHSU, and Stanford is available as part of Release 5.1:
- Level 1-3 and auxiliary 10X Visium data for 18 new participants
- Level 2-3 CODEX data for 20 new participants
- Level 1-2 Electron Microscopy data for 12 new participants
- Level 2 H&E data for 50 new participants
- Level 1 LC-MS/MS data for 7 new participants
- Level 1 LC-MS3 data for 7 new participants
- Level 3-4 MIBI data for 32 new participants
- Level 1, 3, and 4 Mass Spectrometry data for 7 new participants
- Level 1 Shotgun MS (lipidomics) data for 7 new participants
- Level 4 Imaging data for 7 new participants
February 29, 2024
The HTAN DCC has finalized the fifth release of HTAN data. The release includes 9,764 new files for 486 participants and 1,582 biospecimens from several atlases:
-
Level 1-2 data
- 13 participants and 33 biospecimens with 10X Visium
- 216 participants and 614 biospecimens with Bulk DNASeq
- 53 participants and 133 biospecimens with Bulk RNASeq
- 22 participants and 86 biospecimens with CODEX
- 4 participants and 12 biospecimens with CyCIF
- 21 participants and 100 biospecimens with H&E
- 7 participants and 93 biospecimens with LC-MS3
- 12 participants and 26 biospecimens with RPPA
- 3 participants and 5 biospecimens with mIHC
- 151 participants and 369 biospecimens with sc/snRNASeq
- 52 participants and 97 biospecimens with sc/snATACSeq
-
Level 3-4 data
- 42 participants and 63 biospecimens with 10X Visium
- 32 participants and 69 biospecimens with Bulk DNASeq
- 8 participants and 37 biospecimens with CODEX
- 7 participants and 40 biospecimens with CyCIF
- 12 participants and 26 biospecimens with RPPA
- 194 participants and 300 biospecimens with sc/snRNASeq
- 194 participants and 238 biospecimens with sc/snATACSeq
- 8 participants and 37 biospecimens with Imaging
February 9th, 2024
New data from CHOP is available as part of Release 4.2:
- Level 1-2 single nucleus ATAC-seq data for 28 new participants
- Level 1-2 single nucleus RNA-seq data for 30 new participants
- Level 3-4 single nucleus ATAC-seq data for 21 new participants
- Level 3-4 single nucleus RNA-seq data for 23 new participants
- Level 1-3 bulk DNA data for 20 new participants
December 4th, 2023
- 175 additional level 2 imaging files are now available to download through CRDC-GC.
- New feature: Interactive plots tab (beta) on the explore page.
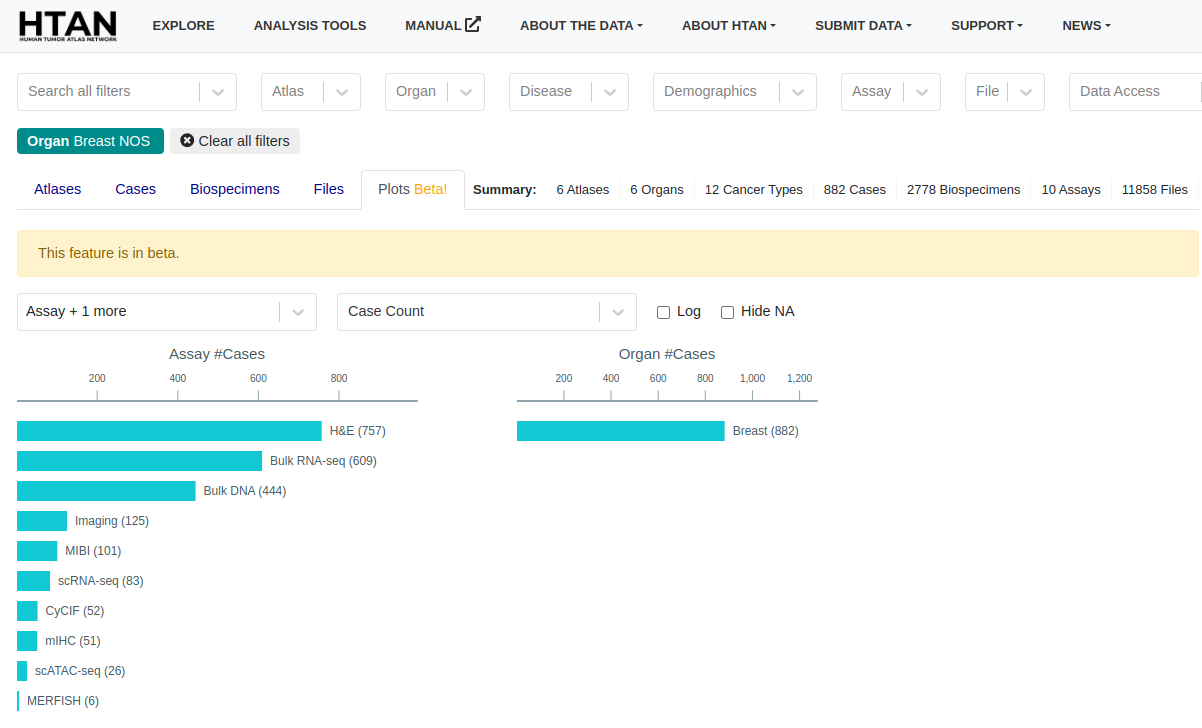
November 20th, 2023
New data from SRRS and BU is available as part of Release 4.1:
- Level 2 H&E images for 9 participants and 72 biospecimens
- Level 2 IMC data for 1 participant and 1 biospecimen
August 25th, 2023
The HTAN DCC has finalized the fourth release of HTAN data. The release includes 13,240 new files for 932 participants and 2,571 biospecimens from several atlases:
-
Level 1-2 data
- 22 participants and 23 biospecimens with 10X Visium
- 352 participants and 419 biospecimens with Bulk DNASeq
- 5 participants and 31 biospecimens with Bulk MethylationSeq
- 506 participants and 596 biospecimens with Bulk RNASeq
- 55 participants and 1056 biospecimens with CyCIF
- 199 participants and 233 biospecimens with H&E
- 4 participants and 27 biospecimens with HI-C-Seq
- 9 participants and 12 biospecimens with MERFISH
- 11 participants and 11 biospecimens with MIBI
- 74 participants and 74 biospecimens with RareCyte Orion
- 53 participants and 269 biospecimens with mIHC
- 98 participants and 111 biospecimens with sc/snRNASeq
- 102 participants and 115 biospecimens with sc/snATACSeq
-
Level 3-4 data
- 22 participants and 23 biospecimens with 10X Visium
- 218 participants and 218 biospecimens with Bulk DNASeq
- 5 participants and 31 biospecimens with Bulk MethylationSeq
- 473 participants and 487 biospecimens with Bulk RNASeq
- 124 participants and 229 biospecimens with sc/snRNASeq
- 100 participants and 112 biospecimens with single nucleus RNA-seq, ATAC-seq and/or multiome data
- 197 participants and 1437 biospecimens with higher level segmentation data for Imaging
-
Minerva Stories with channel selection
- Over 1800 new Minerva Stories
- New Minerva Stories have independent channel selection
- New Minerva Stories have rich metadata descriptions
August 9th, 2023
New data from WUSTL is available as part of Release 3.5:
- Level 1-2 single nucleus ATAC-seq data for 52 new participants
- Level 1-2 single nucleus RNA-seq data for 42 new participants
- Level 3 single nucleus RNA-seq, ATAC-seq and/or multiome data for 43 new participants
April 14th, 2023
Additional level 1/2 data released through dbGaP.
April 12th, 2023
WUSTL Visium data levels have been restructured and additional auxillerary assay files were released.
March 16th, 2023
New data from WUSTL is available as part of Release 3.2:
- Level 2 & 3 single cell RNA-sequencing data from PDAC study
- Level 2 & 3 10X Visium data from PDAC and CCRCC studies
March 10th, 2023
New data is available from WUSTL and MSK atlases as part of Release 3.1:
- Level 1 & 2 10X Visium data from WUSTL PDAC and CCRCC studies
- Level 1-3 single cell and single nucleus RNA-sequencing data from WUSTL PDAC study
- Level 1-4 MSK single cell RNA-seq data
December 21st, 2022
The HTAN DCC has finalized the third release of HTAN data. This v3 release includes new data from several atlases, and includes:
-
Level 1-2 data
- 94 new participants with scRNASeq
- 339 new participants with Bulk DNASeq
- 148 new participants with Bulk RNASeq
- 29 new participants with MIBI
- 5 new participants with H&E
- 2 new participants with CyCIF
-
Level 3-4 data
- 181 new cases with higher level scRNASeq data
- 30 new cases with higher level segmentation data for Imaging
The Level 3-4 data are downloadable via the HTAN Data Portal. The Access-controlled level 1-2 data will be available through Seven Bridges Cancer Genomics Cloud (CGC) in the beginning of next year.
October 26th, 2022
Reintroduce Vanderbilt's cellxgene instance with updated single cell data including cell type annotations.
October 24th, 2022
Remove sequencing data that is not downloadable yet from the explore page. Add a "Download Source" filter button to allow filtering by data repository.
August 30th, 2022
Vanderbilt has Level 3 BulkDNASeq data available now
August 24th, 2022
Publication pages with associated data are now available for:
- Single-cell multiomics reveals increased plasticity, resistant populations, and stem-cell–like blasts in KMT2A-rearranged leukemia (Chen et al, 2022)
- An omic and multidimensional spatial atlas from serial biopsies of an evolving metastatic breast cancer (Johnson et al, 2022)
August 12th, 2022
Imaging data from HMS, OHSU and WashU are now available in DICOM-TIFF format from the Imaging Data Commons (IDC) .
July 11th, 2022
New data available for Trans-Network Project (TNP) SARDANA including CODEX, CyCIF, H&E and SABER.
June 15th, 2022
New WUSTL PDAC data available including 10X Visum, scRNASeq, Bulk DNA and RNA Seq as part of Release 2.3.
June 10th, 2022
All the HTAN MSKCC single cell sequencing data with harmonized cell type information is now available directly from the cellxgene website.
May 27th, 2022
Level 1 and 2 sequencing data are now available through dbGaP.
May 18th, 2022
HTAPP Level 4 scRNA-seq data (CRC) is now available as part of Release 2.2.
March 30th, 2022
Additional HTAN data have been released (Release 2.1).
March 25th, 2022
Disabled Vanderbilt's cellxgene instance pending new data updates.
March 5th, 2022
The second data release of HTAN public data is now available on the HTAN Data Portal. This v2.0 release includes new data from several HTAN atlases:
- 192 cases with new imaging data of which 86 have MIBI data, 105 CyCIF, and 1 both CyCIF and mIHC. This means people can now browse and view over 500 images in Minerva.
- 100 cases with new scRNASeq data of which 62 have level 3 and higher available for download, allowing for download and analysis of scRNASeq data from over 150 cases and 250 biospecimens.
- 1 case with both bulk DNASeq and RNASeq, which can now be explored in cBioPortal as well as downloaded from the data portal.


May 24th, 2021
The first release of HTAN public data is now available on the HTAN Data Portal. This release includes Level 3 and 4 scRNAseq data from several HTAN atlases.